|
With this, we aim to reach the community of non-specialists in proteomics to find a common language and illustrate the basic steps of –omics data processing. This manuscript aims to demonstrate to those who are not familiar with the math and statistics behind these workflows that a proteomics dataset can be processed, simplified and interpreted in software like Microsoft Excel. 
In this paper, we describe key steps of the typical data transformation, normalization and statistics in proteomics data analysis using a simple spreadsheet. The following example shows how to calculate a p-value for a correlation coefficient in Excel. We acknowledge in advance that dedicated scripts and software have a higher level of sophistication but we hereby claim that the approach we describe makes proteomics data processing immediately accessible to every scientist. t r (n-2) / (1-r2) The p-value is calculated as the corresponding two-sided p-value for the t-distribution with n-2 degrees of freedom. This creates a risk or at least a limiting factor, as the biological interpretation of a dataset is contingent of a third-party specialist transforming data without the input of the project leader. However, proteomics has become popular in most other biological and biomedical disciplines, resulting in more and more studies where data processing is delegated to specialists that are not lead authors of the scientific project. Many bioinformatics tools are freely available for the community, some of which within reach for scientists with limited or no background in programming and statistics. A significant component of being a proteomics scientist is the ability to process these tables to identify regulated proteins. 05).Proteomics studies generate tables with thousands of entries. There is a significant difference between the observed and expected genotypic frequencies ( p <. The Χ 2 value is greater than the critical value, so we reject the null hypothesis that the population of offspring have an equal probability of inheriting all possible genotypic combinations. Step 5: Decide whether the reject the null hypothesis The Χ 2 value is greater than the critical value. Note: cant find the Data Analysis button Click here to load the Analysis ToolPak add-in. Step 4: Compare the chi-square value to the critical value On the Data tab, in the Analysis group, click Data Analysis. 05 and df = 3, the Χ 2 critical value is 7.82. P-value is a statistical term that helps you to determine, if the hypothesis you use is true, the probability of the sampling variation. 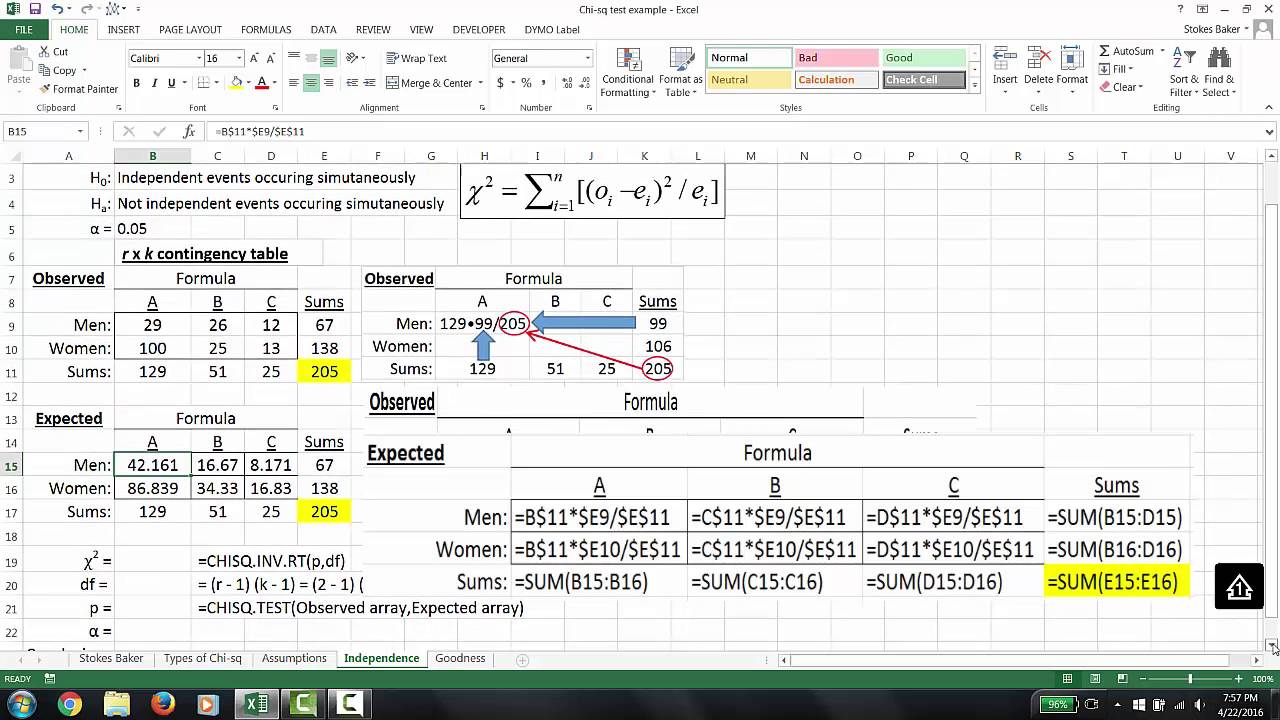
Since there are four groups (round and yellow, round and green, wrinkled and yellow, wrinkled and green), there are three degrees of freedom.įor a test of significance at α =. Χ 2 = 8.41 + 8.67 + 11.6 + 5.4 = 34.08 Step 3: Find the critical chi-square value Below are two methods to calculate a p-value in Excel. The expected phenotypic ratios are therefore 9 round and yellow: 3 round and green: 3 wrinkled and yellow: 1 wrinkled and green.įrom this, you can calculate the expected phenotypic frequencies for 100 peas: Phenotype If the two genes are unlinked, the probability of each genotypic combination is equal. To calculate the expected values, you can make a Punnett square. To find the p-value for this test statistic, we will use the following formula in Excel: T.DIST.RT(2.1689, 14) The following screenshot shows how to use this formula in practice. Step 1: Calculate the expected frequencies This would suggest that the genes are linked. 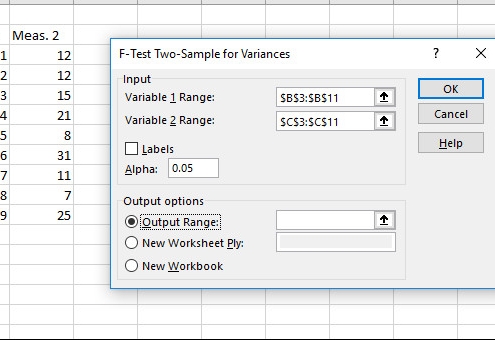
Alternative hypothesis ( H a): The population of offspring do not have an equal probability of inheriting all possible genotypic combinations.This would suggest that the genes are unlinked. To find the p-value that corresponds to a Chi-Square test statistic in Excel, you can use the () function, which uses the following syntax: (x, degfreedom) where: x: The Chi-Square test statistic degfreedom: The degrees of freedom The following examples show how to use this function in practice.Null hypothesis ( H 0): The population of offspring have an equal probability of inheriting all possible genotypic combinations. Introduction to calculating a p-value For a lower-tailed test, the p-value is equal to this probability p-value cdf(ts).The hypotheses you’re testing with your experiment are: You perform a dihybrid cross between two heterozygous ( RY / ry) pea plants. Suppose that you want to know if the genes for pea texture (R = round, r = wrinkled) and color (Y = yellow, y = green) are linked. When genes are linked, the allele inherited for one gene affects the allele inherited for another gene. One common application is to check if two genes are linked (i.e., if the assortment is independent). Chi-square goodness of fit tests are often used in genetics.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
